GapR subunits are differentiated on the basis of color (gray and green), and hydrophobic residues in H2 and H3 are shown as spheres. The upper panel shows a structural representation of the interface involved in GapR tetramerization, derived from the DNA-bound crystal structure (PDB 6CG8) ( 3). (C) Determination of the oligomerization state of WT GapR and two truncated GapR proteins. His 6-HU under the same conditions was used as a control. The molecular weights calculated for a few bands of DNA-bound GapR are shown on the right. His 6-GapR 1–89 purified without nuclease treatment and dialyzed against low-salt buffer was resolved by PAGE under native conditions, and the gel was silver stained. (B) Analysis of DNA-bound GapR by native PAGE. 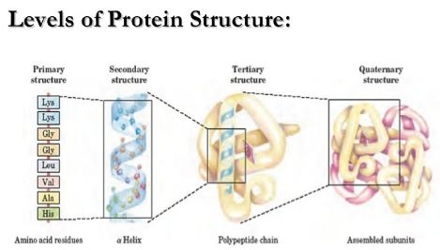
His 6-HU purified without nuclease treatment and dialyzed against high-salt plus EDTA buffer was used as a control. Shown below is a gel used to detect DNA prepared with fractions collected from protein treated under high-salt plus EDTA conditions and stained with ethidium bromide. Fractions 8 to 20 were resolved by SDS-PAGE, and the gels were silver stained. Protein at 50 μM (calculated from the monomeric state) was used for all runs. His 6-GapR 1–89 separated from copurified DNA was also analyzed by size exclusion chromatography (low salt after removal of DNA). His 6-GapR 1–89 purified without nuclease treatment was dialyzed against buffer containing either 150 mM NaCl (low salt) or 1 M NaCl supplemented with 1 mM EDTA (high salt plus EDTA) and analyzed by size exclusion chromatography using the Superdex 200 10/300 GL column. (A) Analysis of the full-length GapR by size exclusion chromatography. This plasticity in the tetramer structure is thought to be important for GapR to translocate along B-DNA, searching for overtwisted regions, and to form a tight complex at overtwisted DNA, where GapR stimulates topoisomerases ( 6).įIG 1 FIG 1 GapR isolated from copurified DNA is a tetramer, but mutant proteins with helix H3 deleted are dimers. Despite the overall folding, the crystal structures slightly deviate from each other with respect to the position of H3, allowing the central channel to accommodate overtwisted or B-DNA ( 6). Moreover, the side chains of positively charged residues at H1 and H2 are pointed toward the central channel of the tetramer ( 3, 6) and are in close proximity to phosphate groups of the encircled DNA ( 3), suggesting their involvement in DNA binding. 
In all these structures, each subunit of GapR folds into three α-helices (H1, H2, and H3), all of them potentially involved in self-association: H1 yielding a dimer, and an interface formed by H2 and H3 promoting the dimer of dimers ( 3, 6). High-resolution crystal structures of DNA-bound GapR revealed a tetrameric protein assembly encircling DNA ( 3, 6).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
